Asymmetric gene expression and cell-type-specific regulatory networks in theroot of bread wheat revealed by single-cell multiomics analysis.
Zhang L, He C, Lai Y, Wang Y, Kang L, Liu A, LanC, Su H, Gao Y, Li Z, Yang F, Li Q, Mao H, Chen D, Chen W, Kaufmann K, Yan W
Genome Biol. 2023 Apr 4;24(1):65. doi: 10.1186/s13059-023-02908-x.
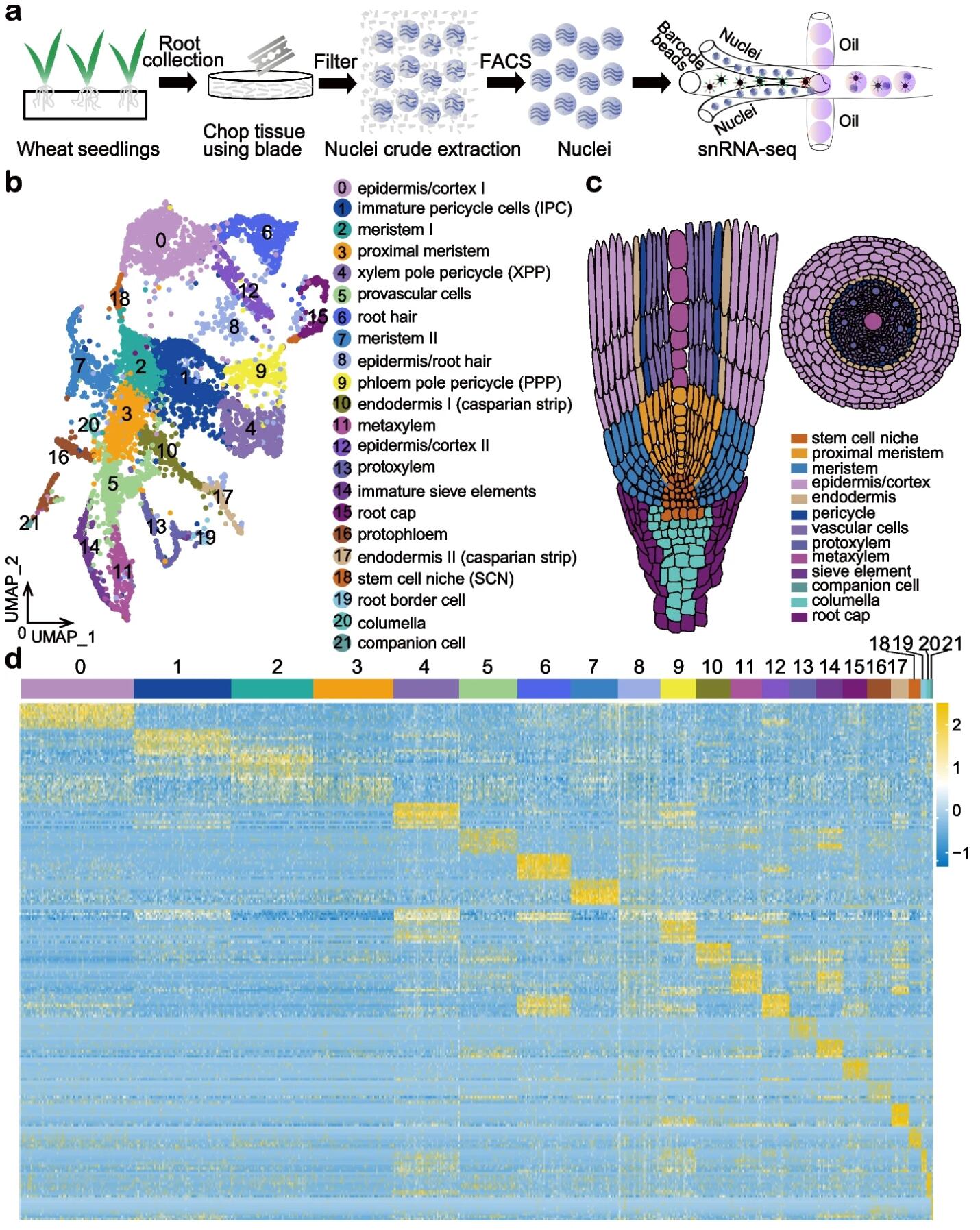
BACKGROUND: Homoeologs are defined as homologous genes resulting fromallopolyploidy. Bread wheat, Triticum aestivum, is an allohexaploid species withmany homoeologs. Homoeolog expression bias, referring to the relativecontribution of homoeologs to the transcriptome, is critical for determining thetraits that influence wheat growth and development. Asymmetric transcription ofhomoeologs has been so far investigated in a tissue or organ-specific manner,which could be misleading due to a mixture of cell types.RESULTS: Here, we perform single nuclei RNA sequencing and ATAC sequencing ofwheat root to study the asymmetric gene transcription, reconstruct celldifferentiation trajectories and cell-type-specific gene regulatory networks. Weidentify 22 cell types. We then reconstruct cell differentiation trajectoriesthat suggest different origins between epidermis/cortex and endodermis,distinguishing bread wheat from Arabidopsis. We show that the ratio ofasymmetrically transcribed triads varies greatly when analyzing at thesingle-cell level. Hub transcription factors determining cell type identity arealso identified. In particular, we demonstrate that TaSPL14 participates invasculature development by regulating the expression of BAM1. Combiningsingle-cell transcription and chromatin accessibility data, we construct thepseudo-time regulatory network driving root hair differentiation. We findMYB3R4, REF6, HDG1, and GATAs as key regulators in this process.CONCLUSIONS: Our findings reveal the transcriptional landscape of rootorganization and asymmetric gene transcription at single-cell resolution inpolyploid wheat.