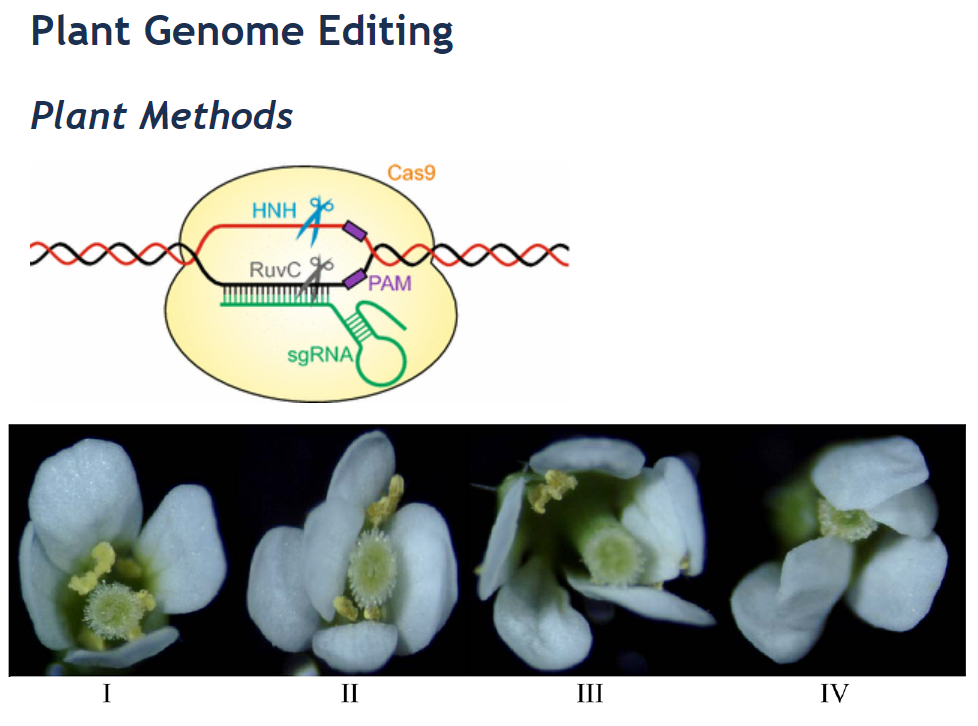
Efficient multiplex mutagenesis by RNA-guided Cas9 and its use in the characterization of regulatory elements in the AGAMOUS gene.
Yan W, Chen D, Kaufmann K
Plant Methods. 2016 Apr 25;12:23. doi: 10.1186/s13007-016-0125-7.
BACKGROUND: The efficiency of multiplex editing in plants by the RNA-guided Cas9 system is limited by efficient introduction of its components into the genome and by their activity. The possibility of introducing large fragment deletions by RNA-guided Cas9 tool provides the potential to study the function of any DNA region of interest in its 'endogenous' environment. RESULTS: Here, an RNA-guided Cas9 system was optimized to enable efficient multiplex editing in Arabidopsis thaliana. We demonstrate the flexibility of our system for knockout of multiple genes, and to generate heritable large-fragment deletions in the genome. As a proof of concept, the function of part of the second intron of the flower development gene AGAMOUS in Arabidopsis was studied by generating a Cas9-free mutant plant line in which part of this intron was removed from the genome. Further analysis revealed that deletion of this intron fragment results 40% decrease of AGAMOUS gene expression without changing the splicing of the gene which indicates that this regulatory region functions as an activator of AGAMOUS gene expression. CONCLUSIONS: Our modified RNA-guided Cas9 system offers a versatile tool for the functional dissection of coding and non-coding DNA sequences in plants.