The genome of hibiscus hamabo reveals its adaptation to saline and waterloggedhabitat.
Wang Z, Xue JY, Hu SY, Zhang F, Yu R, Chen D, Van de PeerY, Jiang J, Song A, Ni L, Hua J, Lu Z, Yu C, YinY, Gu C
Hortic Res. 2022 Mar 23;9:uhac067. doi: 10.1093/hr/uhac067.
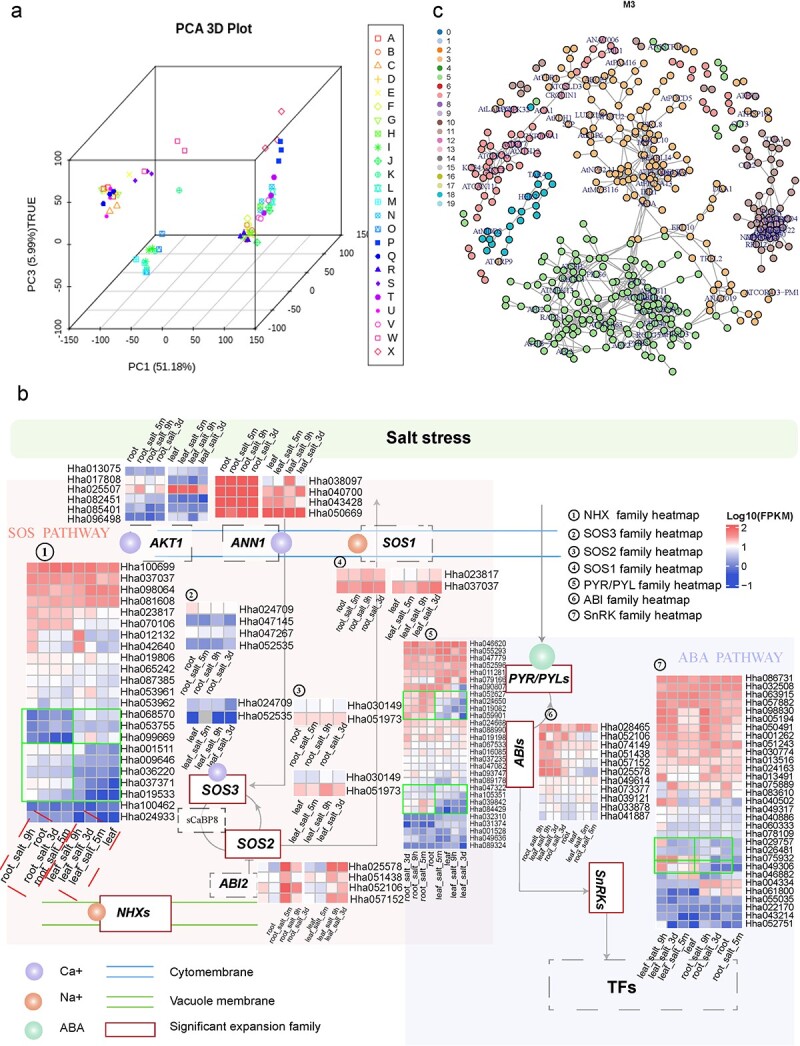
Hibiscus hamabo is a semi-mangrove species with strong tolerance to salt andwaterlogging stress. However, the molecular basis and mechanisms that underliethis strong adaptability to harsh environments remain poorly understood. Here,we assembled a high-quality, chromosome-level genome of this semi-mangrove plantand analyzed its transcriptome under different stress treatments to revealregulatory responses and mechanisms. Our analyses suggested that H. hamabo hasundergone two recent successive polyploidy events, a whole-genome duplicationfollowed by a whole-genome triplication, resulting in an unusually large genenumber (107鈥?09 genes). Comparison of the H. hamabo genome with that of itsclose relative Hibiscus cannabinus, which has not experienced a recent WGT,indicated that genes associated with high stress resistance have beenpreferentially preserved in the H. hamabo genome, suggesting an underlyingassociation between polyploidy and stronger stress resistance. Transcriptomicdata indicated that genes in the roots and leaves responded differently tostress. In roots, genes that regulate ion channels involved in biosynthetic andmetabolic processes responded quickly to adjust the ion concentration andprovide metabolic products to protect root cells, whereas no such rapid responsewas observed from genes in leaves. Using co-expression networks, potentialstress resistance genes were identified for use in future functionalinvestigations. The genome sequence, along with several transcriptome datasets,provide insights into genome evolution and the mechanism of salt andwaterlogging tolerance in H. hamabo, suggesting the importance ofpolyploidization for environmental adaptation.