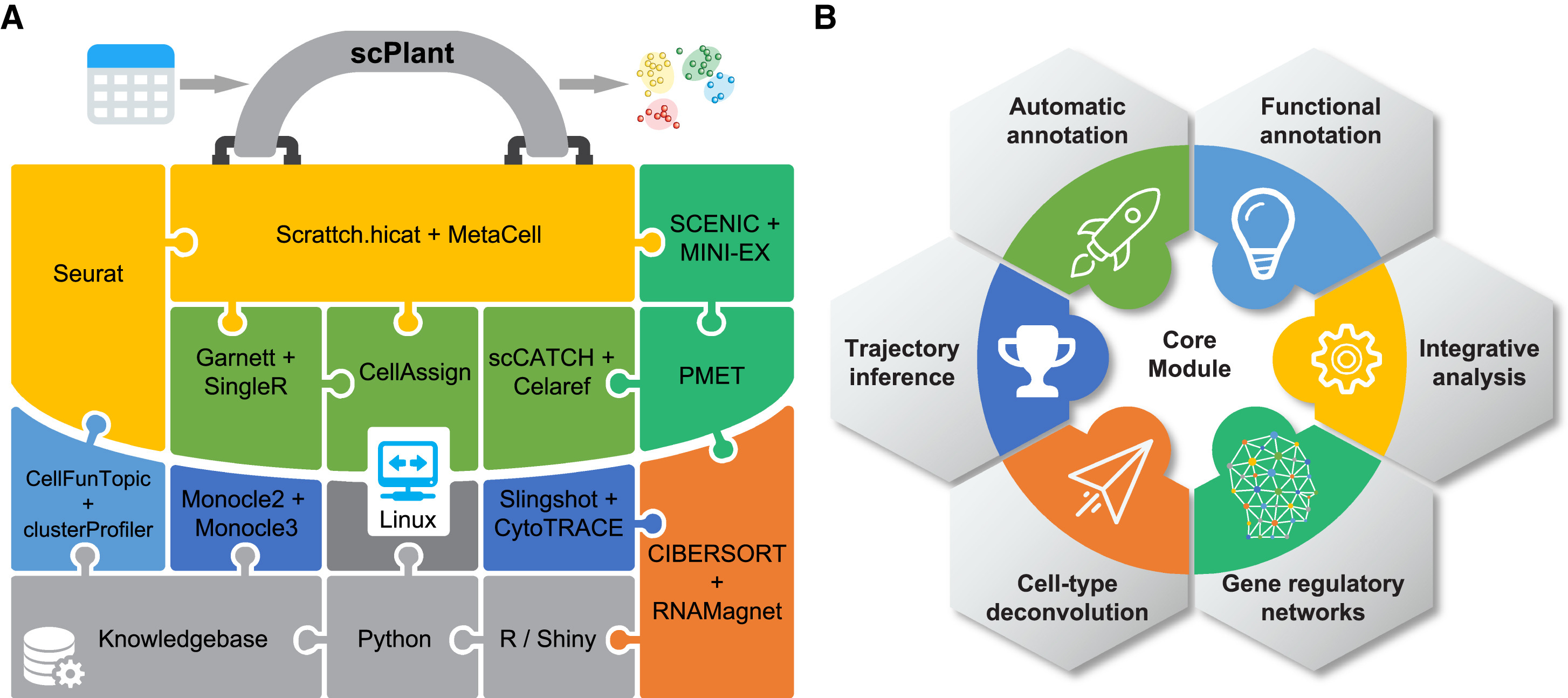
scPlant: A versatile framework for single-cell transcriptomic data analysis in plants.
Cao S, He Z, Chen R, Luo Y, Fu LY, Zhou X, He C, Yan W,Zhang CY, Chen D*
Plant Commun. 2023 May 29:100631. doi: 10.1016/j.xplc.2023.100631.
Single-cell transcriptomics has been fully embraced in plant biological researchand is revolutionizing our understanding of plant growth, development, andresponses to external stimuli. However, single-cell transcriptomic data analysisin plants is not trivial, given that there is currently no end-to-end solutionand that integration of various bioinformatics tools involves a large number ofrequired dependencies. Here, we present scPlant, a versatile framework forexploring plant single-cell atlases with minimum input data provided by users.The scPlant pipeline is implemented with numerous functions for diverseanalytical tasks, ranging from basic data processing to advanced demands such ascell-type annotation and deconvolution, trajectory inference, cross-species dataintegration, and cell-type-specific gene regulatory network construction. Inaddition, a variety of visualization tools are bundled in a built-in Shinyapplication, enabling exploration of single-cell transcriptomic data on the fly.